The consumption of antidepressant medications has skyrocketed in recent decades, reaching more than 337 million prescriptions written in 2016 in the United States alone (Wang et. al. 2019). For many individuals, these drugs are critical to maintaining everyday health as they treat many life-threatening psychiatric disorders. While their exact mechanisms differ, these medications travel in the bloodstream to the brain where they are able to influence the release of chemicals known as neurotransmitters that generate emotional states. However, while their intended target is the brain, these drugs continue to circulate throughout the body, thereby interacting with other organs and structures (Wang et. al. 2019).
In their 2019 study, researchers led by Iva Lukic used data indicating the presence of antidepressants in the digestive tract to investigate the effect of these medications on the gut microbiome. After treating mice with different types of antidepressants, the team noticed a change in the types of bacteria present within the gut when compared to controls (Lukic et. al. 2019). This discovery that antidepressants could impact the types of bacteria present within the body ultimately led researcher Jianhua Guo to question the additional effects that these medications could have on bacteria. As antibiotics have also been shown to affect the composition of the gut microbiome, Guo began by investigating if the antidepressant fluoxetine could help Escherichia coli cells survive in the presence of various antibiotics. After finding that exposure to this medication did increase E. coli’s resistance to antibiotic treatments, Guo decided to expand his hypothesis to examine the overall connection of antidepressant usage with antibiotic resistance in bacteria.
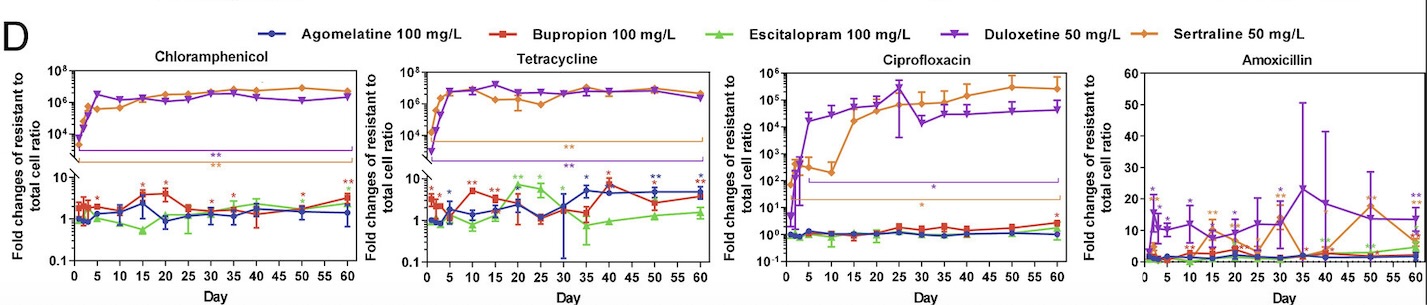
Collaborating with researchers Zue Wang and Zhigang Yue, Guo’s lab began by choosing five major types of antidepressant medications: sertraline, escitalopram, bupropion, duloxetine, and agomelatine. These medications differ in the ways that they prevent the reuptake of serotonin and norepinephrine in the brain, thereby allowing the researchers to examine the effects of various types of antidepressants that may be prescribed to patients. Then, E. coli bacteria were added to media containing varying concentrations of these five antidepressants. Once these cells were treated with antidepressants, the researchers began to test the cells’ resistance against antibiotics. In order to accurately reflect antibiotic use in the real world, the tested antibiotics covered the six main categories of antibiotic medications available on the market. The antidepressant-treated bacteria were then swabbed onto plates containing one of the tested antibiotics to observe cell growth. Based on the growth present on these plates, the researchers were able to estimate the incidence rate of bacterial resistance of E. coli bacteria treated with different antidepressants.
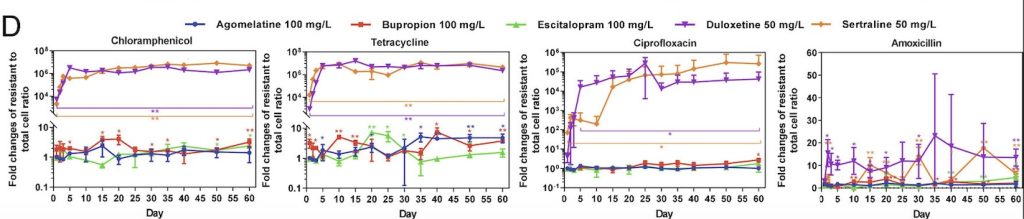
Through this experiment, the lab observed that E. coli cells grown in sertraline and duloxetine, two antidepressants that inhibit the reuptake of serotonin, exhibited the greatest number of resistant cells across all the tested antibiotics (fig. 1). They also noted that E. coli cells exhibiting resistance to one antibiotic often demonstrate some level of resistance to other antibiotics as well. After detecting a correlation between antibiotic-resistance development and exposure to antidepressants, the lab tested the concentration dependence of this effect. While lowering the concentration of antidepressants seemed to decrease the amount of resistant E. coli cells, resistant cells continued to appear on the plates over time, suggesting that lowering antidepressant dosages only prolongs the process of developing antibiotic resistance.
After analyzing this data, the researchers were confronted with a question: what about anti-depressants led to the development of antibiotic resistance in bacteria? To examine this question, the lab used flow cytometry to examine what was happening within the bacterial cells. This lab technique uses a fluorescent dye that binds to specific intercellular target molecules, thereby allowing these components to be visualized. After applying this dye to resistant cells grown on the antibiotic agar plates, the researchers noticed the presence of specific oxygen compounds known as reactive oxygen species (ROS). Unstable ROS bind to other molecules within a cell, disrupting normal functioning and causing stress. Elevated cellular stress levels have been shown to induce the transcription of specific genes in bacteria that produce proteins to help return the cell to normal functioning (Wang et. al. 2019).
ROS molecules have been shown to induce the production of efflux pumps in bacteria, leading the lab to investigate if these structures were involved in the antibiotic resistance of E. coli cells. Efflux pumps are structures in the cell membrane of a protein that pump harmful substances out of the cell. The lab mapped the genome to look for activated genes associated with the production of this protein. According to the computer model, more DNA regions in resistance bacteria coding for efflux pumps were active than in the Wild Type. The researchers then concluded that efflux pumps were being produced in response to antidepressant exposure. These additional efflux pumps removed antibiotic molecules in resistant E. coli, thereby allowing them to survive in the presence of lethal drugs.
The antibiotic resistance uncovered in this study was significant and persistent. Even one day of exposure to antidepressants like sertraline and duloxetine led to the presence of resistant cells. Furthermore, the team demonstrated that these antibiotic-resistant capabilities often do not disappear over time; rather, they are inherited between generations of bacteria, leading to the proliferation of dangerous cells unsusceptible to available treatments. The next logical step towards validating the connection between antidepressants and antibiotic resistance would include studying the gut microbiomes of patients taking anti-depressants to look for antibiotic-resistant bacteria.
This study reveals a novel issue that must be attended to. In 2019, 1.27 million deaths worldwide could be directly attributed to antibiotic-resistant microbes, a number expected to grow to 10 million by the year 2050 (O’Neill 2023). These “superbugs” present a dangerously growing reality. If the correlation between antidepressant use and antibiotic resistance is left uninvestigated, superbugs will likely continue to develop even as antibiotic use is regulated and monitored to battle them. Only by taking this connection seriously will researchers be able to fully grapple with and battle the growing antibiotic resistance trends, thereby preventing common infections from becoming death sentences.
Sources:
CDC. (2022, July 15). The biggest antibiotic-resistant threats in the U.S. Centers for Disease Control and Prevention. https://www.cdc.gov/drugresistance/biggest-threats.html
Drew, L. (2023). How antidepressants help bacteria resist antibiotics. Nature. https://doi.org/10.1038/d41586-023-00186-y
Jin, M., Lu, J., Chen, Z., Nguyen, S. H., Mao, L., Li, J., Yuan, Z., & Guo, J. (2018). Antidepressant fluoxetine induces multiple antibiotics resistance in Escherichia coli via ROS-mediated mutagenesis. Environment International, 120, 421–430. https://doi.org/10.1016/j.envint.2018.07.046
Lukić, I., Getselter, D., Ziv, O., Oron, O., Reuveni, E., Koren, O., & Elliott, E. (2019). Antidepressants affect gut microbiota and Ruminococcus flavefaciens is able to abolish their effects on depressive-like behavior. Translational Psychiatry, 9(1), 1–16. https://doi.org/10.1038/s41398-019-0466-x
O’Neill, J. (Ed.). (2016). Tackling Drug-Resistant Infections Globally: Final Report and Recommendations. The Review on Antimicrobial Resistance. https://amr-review.org/sites/default/files/160518_Final%20paper_with%20cover.pdf
Thompson, T. (2022). The staggering death toll of drug-resistant bacteria. Nature. https://doi.org/10.1038/d41586-022-00228-x
Wang, Y., Yu, Z., Ding, P., Lu, J., Mao, L., Ngiam, L., Yuan, Z., Engelstädter, J., Schembri, M. A., & Guo, J. (2023). Antidepressants can induce mutation and enhance persistence toward multiple antibiotics. Proceedings of the National Academy of Sciences, 120(5), e2208344120. https://doi.org/10.1073/pnas.2208344120