mRNA vaccines have emerged from the COVID-19 pandemic as a promising approach to fighting infectious diseases (Kutikuppala et al. 2024). Different from traditional vaccines, which use a weakened version of a virus, mRNA vaccines deliver an mRNA corresponding to a protein on the surface of the virus. This mRNA allows our immune system to recognize and make small parts of the virus so that it can create antibodies to combat it (Cleveland Clinic 2024). However, mRNA is an unstable and negatively charged molecule, so it must be encased with some type of transport protection to prevent its degradation during delivery (Kutikuppala et al. 2024).
Lipid nanoparticles, or LNPs, have gained popularity in recent years as an effective delivery platform for mRNA vaccines due to their highly tunable composition and their ability to prevent nucleic acid degradation (Xu et al., 2025). One popular example is the utilization of LNPs in the Moderna mRNA-1273 COVID vaccine, where mRNA encoding the protein on the outside of the virus that is recognized by the immune system was encapsulated in an LNP. Vaccination with this LNP-encapsulated mRNA resulted in 90% lower risk of contracting COVID within 21 days for those over the age of 16, demonstrating the power and possibility of this technology (Noor, 2021).
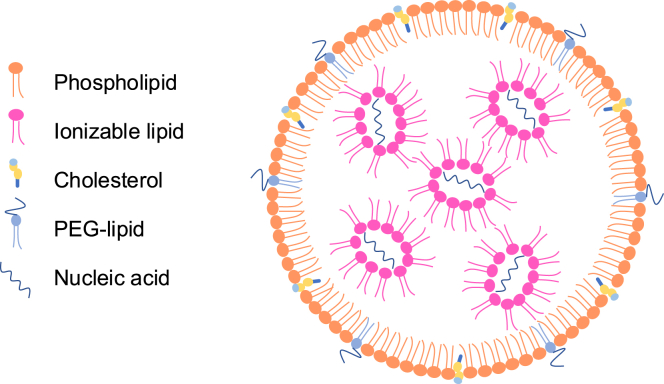
LNPs are extremely small particles composed of: 1) ionizable lipids, which act as a case for the nucleic acid being delivered; 2) phospholipids regulating cell membrane fusion; 3) PEG-lipids and 4) cholesterol which both affect its size and stability (Figure 1) (Xu et al., 2025). An LNP’s formulation can have substantial effects on its ability to avoid cellular barriers for vaccine mRNA entry to a targeted area. For example, degradation of LNPs by enzymes and/or other immune cells after entering the body can affect a vaccine’s ability to reach the targeted tissue (Hou et al., 2021). This is especially critical for scientists working on immunotherapies, as a variation in lipid composition can determine whether the LNP will be taken up by immune cells like dendritic cells or other antigen-presenting cells, which present the LNP to other immune cells and start the immune response (Hou et al., 2021).
Hydrogels have also been utilized by scientists as vaccine carriers that can also augment immune responses. Hydrogels are natural or synthetic materials containing a 3D network of cross-linked polymer chains that allow them to absorb large amounts of a target substance (Ho et al. 2022). Depending on the composition of the hydrogel, scientists have found evidence of greatly increased immune cell recruitment and prolonged immune memory in mouse models of melanoma after a hydrogel-based vaccine was delivered (Kerr et al., 2023; Pal et al., 2024). In other words, the immune response was stronger and also more effective upon encountering a pathogen a second time. Therefore, if a hydrogel were to be used to deliver an LNP, finding the right composition is extremely important, as it can greatly impact its efficacy.
In their paper, Zhu et al. report the effectiveness of LiNx, a nanofiber-hydrogel composite (NHC) mRNA LNP matrix, in tumor and melanoma mouse models. Essentially, they embedded their LNPs within the 3D network of extremely small and cross-linked fibers in a hydrogel to significantly boost the immune response to cancer.
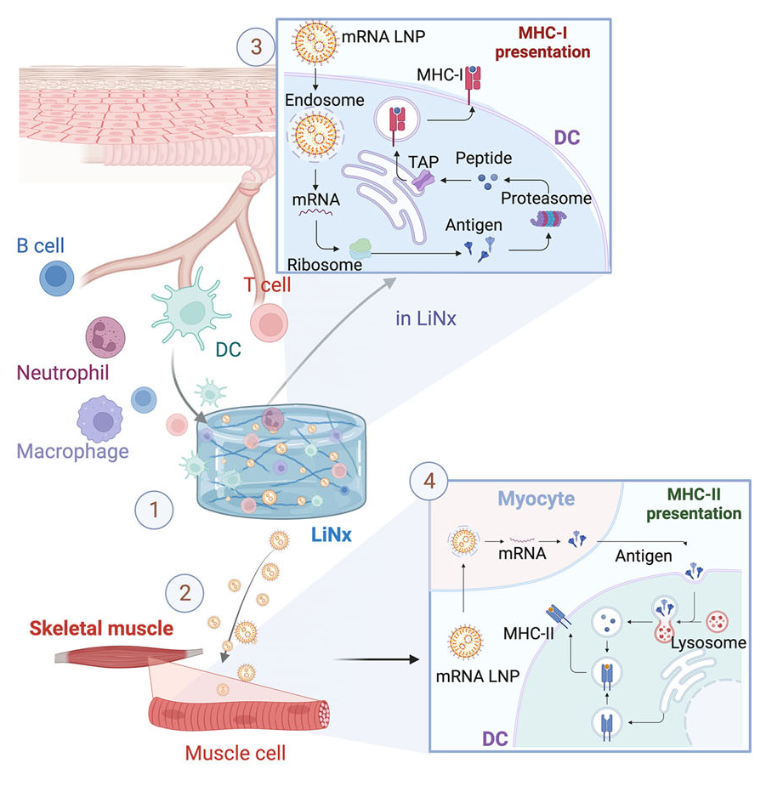
LiNx works as a subcutaneous injection combining the potent immune activation capability of LNPs with the immunostimulatory microenvironment provided by the NHC. While the NHC recruits immune cells to the injection site and promotes immune cell signaling, the LNPs introduce nearby cells to the encapsulated mRNA, resulting in a coordinated adaptive immune response (Figure 2).
The lipid composition of LNPs can affect not only their size and stability, but also their transfection and delivery efficacy, or their ability to deliver the vaccine mRNA into host cells like dendritic cells (which start the immune response). To identify the top-performing LNP formulations, the researchers screened over one thousand different lipid compositions. Three top-performing LNP formulations were identified based on their transfection efficiency in bone marrow-derived dendritic cells: C10, D6, and F5. All of these formulations also separately activated powerful Th1 responses, a type of immune response meant to eliminate bacteria, viruses, and cancer cells, after three doses of subcutaneous injections.
To simply quantify the host cell recruitment and transfection profile of the three different formulations, the researchers injected LiNx containing C10, D6, or F5 LNP into mice and measured the present immune cells 3 and 7 days post-injection. At both 3 and 7 days post-injection, a considerable amount of host cells were found in the NHC scaffold for all three formulations. The D6 formulation showed the greatest host cell recruitment, having a 12.6-fold increase compared to the control.
The researchers then performed a similar experiment, injecting mice with LiNx loaded with a test mRNA to get a better idea of the performance of each formulation compared to each other. They found that 10 days after injection, the D6 formulation contained over one-hundred-fold more transfected cells than C10 and F5-mRNA LiNx. Fourteen days post-injection, the D6-mRNA LiNx was also found to have recruited a more diverse range of immune cells associated with robust and specific immune responses like T cells and B cells. On the other hand, the C10 and F5-mRNA LiNx recruited more immune cells associated with general immune responses, like neutrophils. This shows that the D6-mRNA LiNx induces a stronger and more customized immune response. Additionally, three months post-vaccination, there were 10x more central memory T cells present in the spleens of D6-mRNA mice compared to the control and other formulations, indicating a stronger long-term memory response. These results suggest that the D6 LiNx is the most efficient LiNx formulation.
Having characterized the immune activation induced by D6-mRNA LiNx, the researchers then tested its effectiveness in cancer mouse models. Mice were inoculated with colorectal cancer cells and received vaccinations of one of the LiNx formulations four days later. These mice were administered the vaccines in a single dose, while a separate control group received three doses of only D6 LNPs. The negative control group received only the NHC and protein without the LNP. The median survival time of the single-dose D6 LiNx mice was 75 days compared to 31 days for the negative control group and 37.5 days for the three-dose group, underscoring a heightened tumor suppression response. Fifty percent of these mice remained tumor-free after 100 days. This experiment demonstrated LiNx’s anti-cancer potential in vivo.
In their paper, Zhu et al. demonstrated the effectiveness of a dual-modal approach to cancer immunotherapy. Through the combination of lipid nanoparticle mRNA delivery and a hydrogel microenvironment, they were able to induce a substantially stronger immune response characterized by tumor suppression and long-term immune memory in mouse models. The superior performance of a singular dose of D6 LiNx compared to three LNP doses illustrates the promise found in combining delivery methods with immune-boosting materials for the future development of stronger and longer-lasting cancer immunotherapies.
References:
Ho T-C et al. 2022. Hydrogels: Properties and Applications in Biomedicine. Molecules. 27(9):2902.
Hou X, Zaks T, Langer R, Dong Y. 2021. Lipid nanoparticles for mRNA delivery. Nat Rev Mater. 6(12):1078–1094.
Kerr MD et al. 2023. Biodegradable scaffolds for enhancing vaccine delivery. Bioeng Transl Med. 8(6):e10591.
Kutikuppala LVS et al. 2024. Prospects and Challenges in Developing mRNA Vaccines for Infectious Diseases and Oncogenic Viruses. Med Sci (Basel). 12(2):28.
mRNA Vaccines: What They Are & How They Work. 2024. Cleveland Clinic; [accessed 2026 May 2]. https://my.clevelandclinic.org/health/treatments/21898-mrna-vaccines
Noor R. 2021. Developmental Status of the Potential Vaccines for the Mitigation of the COVID-19 Pandemic and a Focus on the Effectiveness of the Pfizer-BioNTech and Moderna mRNA Vaccines. Curr Clin Microbiol Rep. 8(3):178–185.
Pal S et al. 2024. Extracellular Matrix Scaffold-Assisted Tumor Vaccines Induce Tumor Regression and Long-Term Immune Memory. Adv Mater. 36(15):e2309843.
Xu S et al. 2025. Lipid nanoparticles: Composition, formulation, and application. Mol Ther Methods Clin Dev. 33(2):101463.
Zhu Y et al. 2025. An mRNA lipid nanoparticle-incorporated nanofiber-hydrogel composite for cancer immunotherapy. Nat Commun. 16(1):5707.